 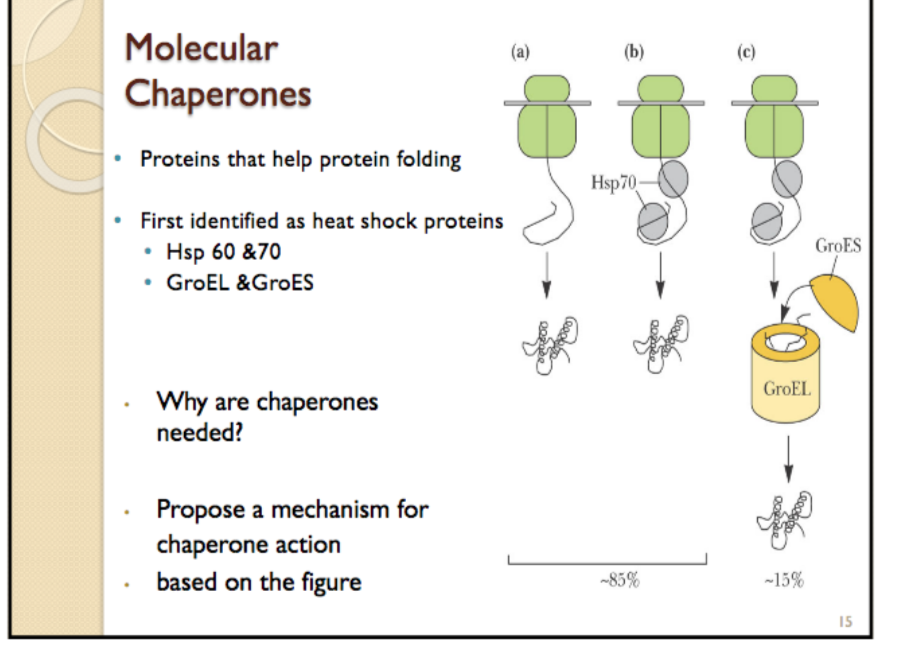
A number of models of chaperonin action have been proposed, which generally focus on two (not mutually exclusive) roles of chaperonin interior: passive and active. According to recent analyses by different experimental techniques, GroEL-bound substrate proteins populate an ensemble of compact and locally expanded states that lack stable tertiary interactions. The exact mechanism by which chaperonins facilitate folding of substrate proteins is unknown. Upon release, the substrate protein will either be folded or will require further rounds of folding, in which case it can again be bound by a chaperonin. These conformational changes allow the chaperonin to bind an unfolded or misfolded protein, encapsulate that protein within one of the cavities formed by the two rings, and release the protein back into solution. Mechanism of action Ĭhaperonins undergo large conformational changes during a folding reaction as a function of the enzymatic hydrolysis of ATP as well as binding of substrate proteins and cochaperonins, such as GroES. It might represent another ancient type of chaperonin. Ī Group I chaperonin gp146 from phage EL does not use a lid, and its donut interface is more similar to Group II. They are thought to be an ancient relative of Group II. They have a lid, but the lid opening is noncooperative in them. Group III includes some bacterial Cpns that are related to Group II. They also interact with a co-chaperone, prefoldin, that helps move the substrate in. They instead contain a "built-in" lid that closes in an ATP-dependent manner to encapsulate its substrates, a process that is required for optimal protein folding activity. Group II chaperonins are not thought to utilize a GroES-type cofactor to fold their substrates. It has been shown to fold the mitochondrial protein rhodanese however, no natural substrates have yet been identified. Methanococcus maripaludis chaperonin (Mm cpn) is composed of sixteen identical subunits (eight per ring). TRiC was originally thought to fold only the cytoskeletal proteins actin and tubulin but is now known to fold dozens of substrates. TRiC, the eukaryotic chaperonin, is composed of two rings of eight different though related subunits, each thought to be represented once per eight-membered ring.A homo-16mer in some archaea, it is regarded as the prototypical type II chaperonin. The complex in archaea is called the thermosome.Group II chaperonins (TCP-1), found in the eukaryotic cytosol and in archaea, are more poorly characterized. Structure of Saccharomyces cerevisiae TRiC in the AMP-PNP bound state ( PDB: 5GW5). Some bacteria use multiple copies of this chaperonin, probably for different peptides. The crystal structure of Escherichia coli GroEL has been resolved to 2.8 Å. The RuBisCO subunit binding protein is a member of this family. The binding of cpn10 to cpn60 inhibits the weak ATPase activity of cpn60. The cpn10 and cpn60 oligomers also require Mg 2+-ATP in order to interact to form a functional complex. The Cpn60 subfamily was discovered in 1998. GroEL/GroES may not be able to undo protein aggregates, but kinetically it competes in the pathway of misfolding and aggregation, thereby preventing aggregate formation. It is like a cover that covers GroEL (box/bottle). GroES (is a single-ring heptamer that binds to GroEL in the presence of ATP or transition state analogues of ATP hydrolysis, such as ADP-AlF 3.GroEL is a double-ring 14mer with a greasy hydrophobic patch at its opening and can accommodate the native folding of substrates 15-60 kDa in size. 
coli is a Group I chaperonin and the best characterized large (~ 1 MDa) chaperonin complex. Group I chaperonins (Cpn60) are found in bacteria as well as organelles of endosymbiotic origin: chloroplasts and mitochondria. Chaperonin proteins may also tag misfolded proteins to be degraded. Molecular chaperones catalyze protein refolding by accelerating partial unfolding of misfolded proteins, aided by energy supplied by the hydrolysis of adenosine triphosphate (ATP). Most proteins spontaneously fold into their most stable three-dimensional conformation, which is usually also their functional conformation, but occasionally proteins mis-fold. The energy to fold proteins is supplied by non-covalent interactions between the amino acid side chains of each protein, and by solvent effects. Newly made proteins usually must fold from a linear chain of amino acids into a three-dimensional tertiary structure. HSP60 belong to a large class of molecules that assist protein folding, called molecular chaperones. They prevent misfolding of proteins during stressful situations such as high heat, by assisting protein folding. 
HSP60, also known as chaperonins ( Cpn), is a family of heat shock proteins originally sorted by their 60kDa molecular mass. Structure of the bacterial chaperonin GroEL. 
0 Comments
Leave a Reply. |



 RSS Feed
RSS Feed
